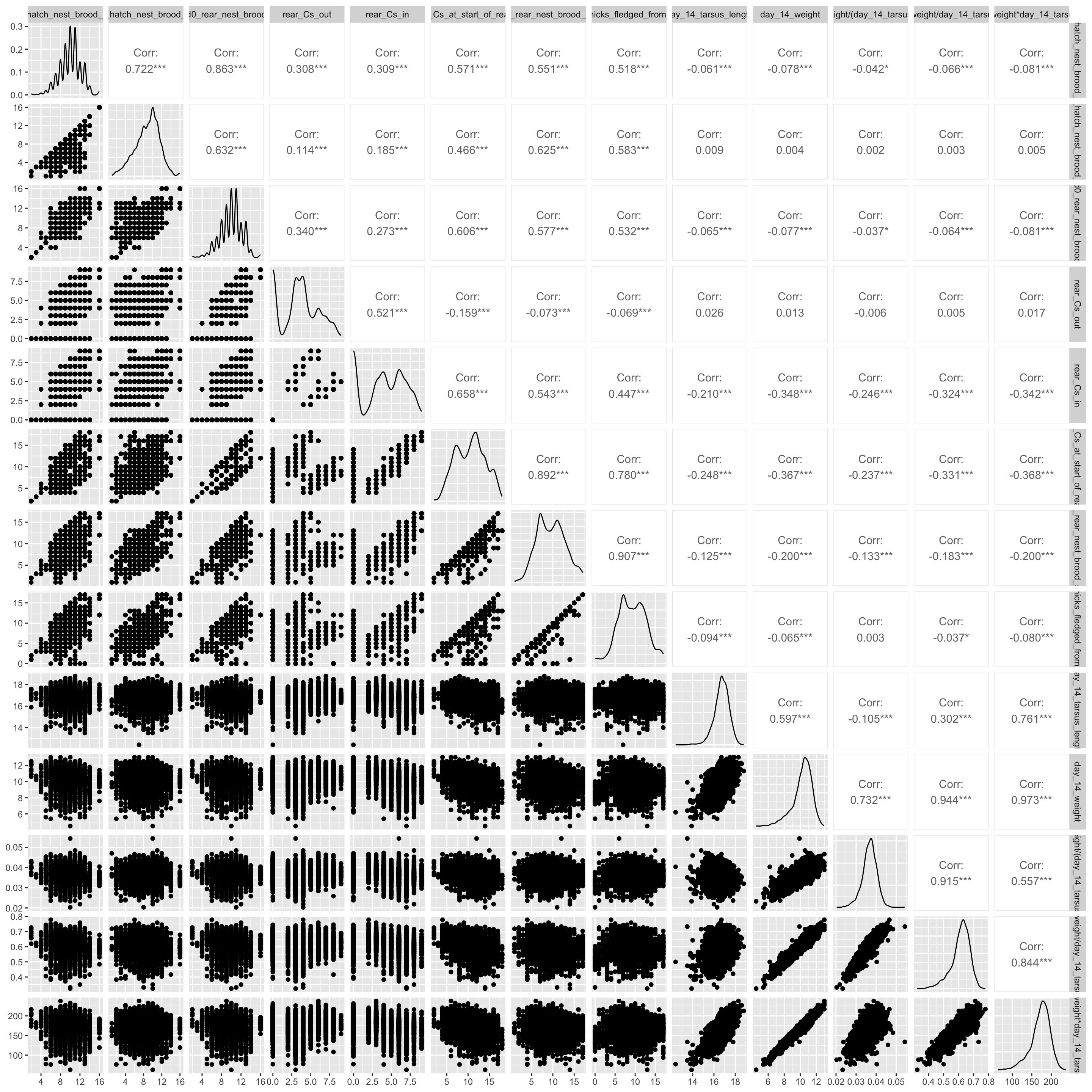
SM D: Correlation Matrices of Case Study Data
Pairwise-correlation plots for the Eucalyptus and blue tit case-study data provided to analysts are shown in Figure D.1 and Figure D.2, respectively. Plots were created with R package GGally (Schloerke et al. 2024).
Code

Date, Quadrat no, Easting, Northing.
Code
blue_tit_data %>%
naniar::replace_with_na_all(condition = ~ .x == ".") %>%
mutate(across(c(contains("_ring"),
rear_nest_trt,
hatch_year,
hatch_nest_breed_ID,
hatch_Area,
hatch_Box,
day14_measurer,
contains("hatch_Box"),
starts_with("rear_"),
starts_with("hatch_nest"),
home_or_away,
-rear_d0_rear_nest_brood_size,
contains("manipulation"),
chick_sex_molec,
Date_of_day14,
`Extra-pair_paternity`,
-rear_Cs_in,
-rear_Cs_out,
chick_survival_to_first_breed_season,
-rear_Cs_at_start_of_rearing),
as.factor),
across(where(is.character), as.numeric),
across(c(rear_Cs_out,
rear_Cs_in,
rear_Cs_at_start_of_rearing),
as.integer)) %>%
select(where(is.numeric), -`day 14 weight`) %>%
GGally::ggpairs()